Building Cython code#
Cython code must, unlike Python, be compiled. This happens in two stages:
A
.pyxor.pyfile is compiled by Cython to a.cfile, containing the code of a Python extension module.The
.cfile is compiled by a C compiler to a.sofile (or.pydon Windows) which can beimport-ed directly into a Python session. setuptools takes care of this part. Although Cython can call them for you in certain cases.
To understand fully the Cython + setuptools build process, one may want to read more about distributing Python modules.
There are several ways to build Cython code:
Write a setuptools
setup.py. This is the normal and recommended way.Run the
cythonizecommand-line utility. This is a good approach for compiling a single Cython source file directly to an extension. A source file can be built “in place” (so that the extension module is created next to the source file, ready to be imported) withcythonize -i filename.pyx.Use Pyximport, importing Cython
.pyxfiles as if they were.pyfiles (using setuptools to compile and build in the background). This method is easier than writing asetup.py, but is not very flexible. So you’ll need to write asetup.pyif, for example, you need certain compilations options.Run the
cythoncommand-line utility manually to produce the.cfile from the.pyxfile, then manually compiling the.cfile into a shared object library or DLL suitable for import from Python. (These manual steps are mostly for debugging and experimentation.)Use the [Jupyter] notebook or the [Sage] notebook, both of which allow Cython code inline. This is the easiest way to get started writing Cython code and running it.
Currently, using setuptools is the most common way Cython files are built and distributed. The other methods are described in more detail in the Source Files and Compilation section of the reference manual.
Building a Cython module using setuptools#
Imagine a simple “hello world” script in a file hello.pyx:
def say_hello_to(name):
print(f"Hello {name}!")
The following could be a corresponding setup.py script:
from setuptools import setup
from Cython.Build import cythonize
setup(
name='Hello world app',
ext_modules=cythonize("hello.pyx"),
)
To build, run python setup.py build_ext --inplace. Then simply
start a Python session and do from hello import say_hello_to and
use the imported function as you see fit.
Using the Jupyter notebook#
Cython can be used conveniently and interactively from a web browser through the Jupyter notebook. To install Jupyter notebook, e.g. into a virtualenv, use pip:
(venv)$ pip install jupyter
(venv)$ jupyter notebook
To enable support for Cython compilation, install Cython as described in the installation guide
and load the Cython extension from within the Jupyter notebook:
%load_ext Cython
Then, prefix a cell with the %%cython marker to compile it
%%cython
a: cython.int = 0
for i in range(10):
a += i
print(a)
%%cython
cdef int a = 0
for i in range(10):
a += i
print(a)
You can show Cython’s code analysis by passing the --annotate option:
%%cython --annotate
...
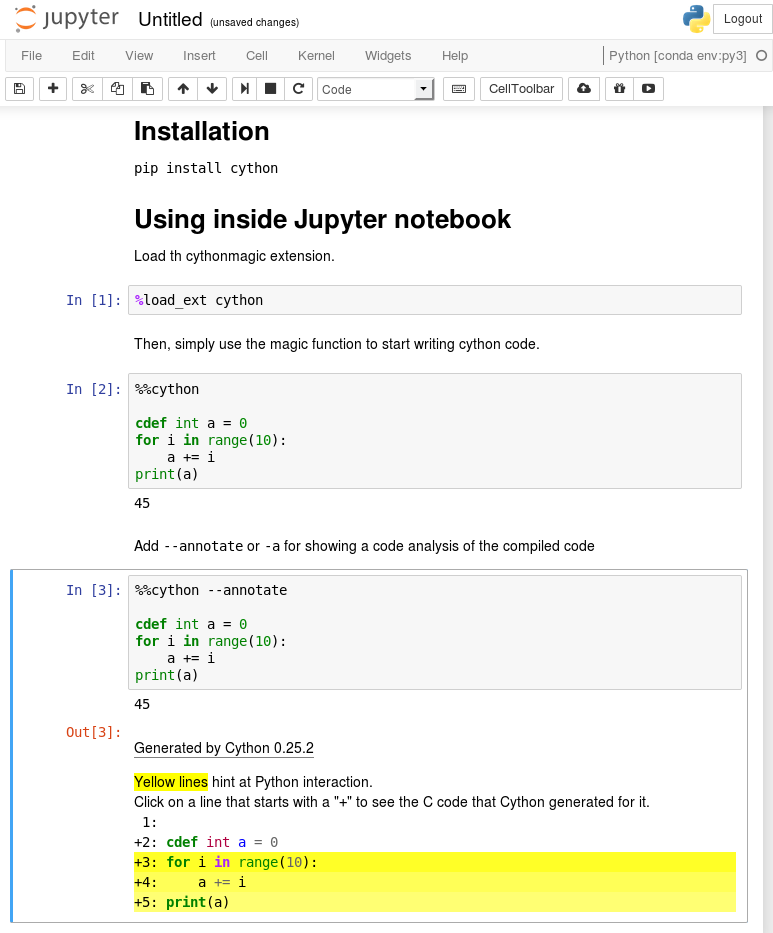
For more information about the arguments of the %%cython magic, see
Compiling with a Jupyter Notebook.
Using the Sage notebook#
For users of the SageMath distribution, the Sage notebook uses Jupyter by default. Sage provides its own implementation of the
%%cythoncell magic (see Sage Magics) which is loaded by default. Alternatively, you can overwrite it with the implementation provided by Cython, the procedure outlined for Jupyter notebooks also applies.

